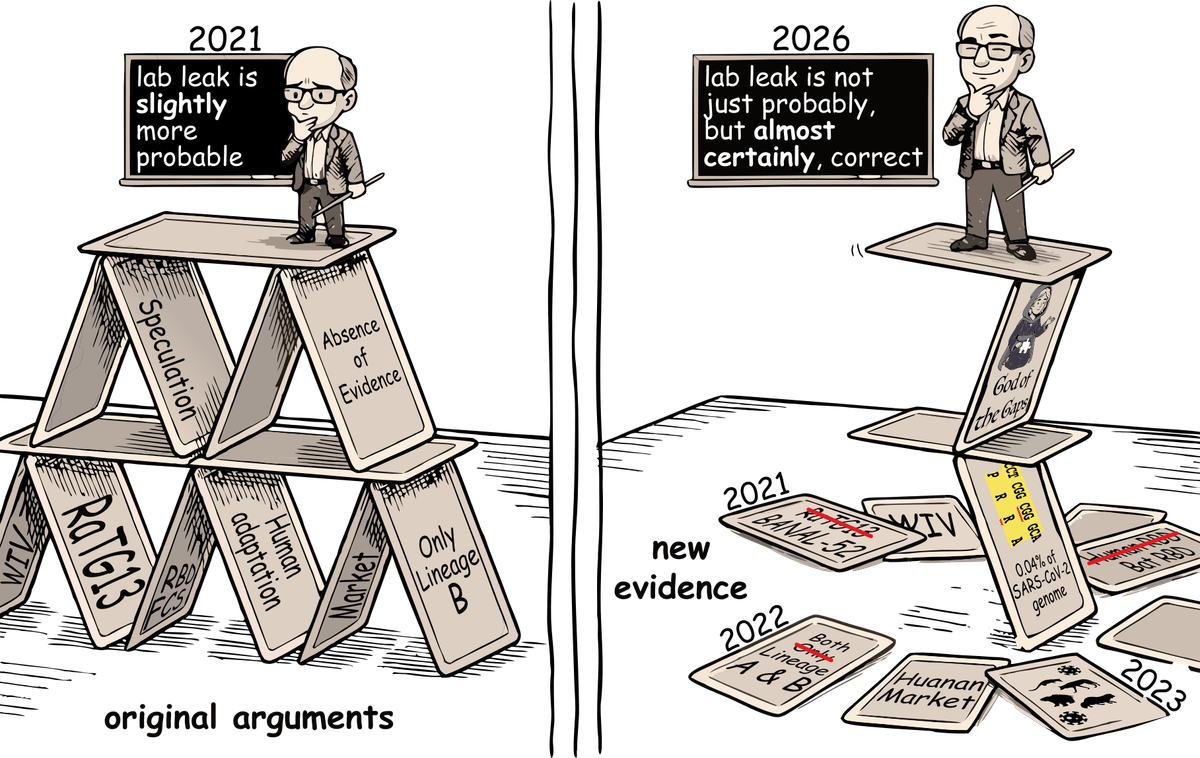
This market resolves once we have a definitive answer to this question. (i.e. "I've looked at all notable evidence presented by both sides and have upwards of 98% confidence that a certain conclusion is correct, and it doesn't seem likely that any further relevant evidence will be forthcoming any time soon.")
This will likely not occur until many years after Covid is no longer a subject of active political contention, motivations for various actors to distort or hide inconvenient evidence have died down, and a scientific consensus has emerged on the subject. For exactly when it will resolve, see /IsaacKing/when-will-the-covid-lab-leak-market
I will be conferring with the community extensively before resolving this market, to ensure I haven't missed anything and aren't being overconfident in one direction or another. As some additional assurance, see /IsaacKing/will-my-resolution-of-the-covid19-l
(For comparison, the level of evidence in favor of anthropogenic climate change would be sufficient, despite the existence of a few doubts here and there.)
If we never reach a point where I can safely be that confident either way, it'll remain open indefinitely. (And Manifold lends you your mana back after a few months, so this doesn't negatively impact you.)
"Come from a laboratory" includes both an accidental lab leak and an intentional release. It also counts if COVID was found in the wild, taken to a lab for study, and then escaped from that lab without any modification. It just needs to have actually been "in the lab" in a meaningful way. A lab worker who was out collecting samples and got contaminated in the wild doesn't count, but it does count if they got contaminated later from a sample that was supposed to be safely contained.
In the event of multiple progenitors, this market resolves YES only if the lab leak was plausibly responsible for the worldwide pandemic. It won't count if the pandemic primarily came from natural sources and then there was also a lab leak that only infected a few people.
I won't bet in this market.
People are also trading
For anyone who finds the argument from Matt Ridley in Viral, recently presented at NIH, compelling, here is a post that might make you reconsider: https://www.the-gallop.com/review-matt-ridleys-viral-case-for-a-lab-leak/
I challenge anyone to find anything from a “zoonosis extremist” as Matt Ridley calls us, that’s as intellectually bankrupt as his edits to Viral between its first and second edition.
The next post on the blog will show how the FBI “lab leak” conclusion in 2021 was based at least partly on copying a stupid reading comprehension error from Twitter conspiracy theorists.
The whistleblower's written testimony is here: https://www.hsgac.senate.gov/wp-content/uploads/letter-and-testimomy.pdf
If you take it at face value, lower level analysts at CIA favored lab leak. Upper management was also listening to a bunch of expert scientists listed by Anthony Fauci. According to the whistleblower, upper management overrode the analysts to make a non-call on origins. Later, under new management (Ratcliffe), on no new information, they changed it to an assessment favoring lab leak.
Erdman faults the IC for not doing what open source sleuths were doing i.e. piece together sparse data to make inferences. He believes that they were doing a better job than the IC. His reddit post features a host of disparate theories, with some clearly inspired by Steven Quay. https://www.reddit.com/r/conspiracy/comments/ni98ax/questions_you_should_ask_about_covid_19/
It's bad news for those like @bens and others who think that there exists secret and unfalsifiable IC evidence that can prove their case.
But most interestingly, he gives a timeline that appears to be based on all of the internal emails that he has had access to as a member of the DIG. In the timeline he states "Credible reporting does indicate that the pandemic may have begun in November 2019 in Wuhan, China and was the product of a lab incident."
This is also the start date for where scientific consensus on origins lies. It is fatal to lab leak which proposes cryptic undetected transmission for several months leading to the establishment of two lineages, the market being a superspreading location. Hence the suspicion based on the purported September database shutdown etc. A November timeline would however require two lab leaks of different lineages that would also have to survive the 4% combined odds of successfully establishing, and subsequent transport to the market without any sign of being detected elsewhere in Wuhan.
@BW Elsewhere Washburne who spent some time in the govt claims that he has had access to non-public information, chatted with the MI6 director etc. One of the claims he makes is that Biden and Blinken collaborated with Xi to shut down the disclosures required under the Covid origins act and berates the author for talking about conspiracies. https://open.substack.com/pub/drbobmorris/p/how-to-build-a-conspiracy-theory?r=g2g7h&utm_campaign=comment-list-share-cta&utm_medium=web&comments=true&commentId=257563246
Erdman's first gateway drug to COVID-19 origins conspiracy theories was the silliness from Sørensen and Dalgleish (which is in the SARS2-engineered-from-HIV genre of conspiracy theories). Quay was next, which is why Erdman was all about Orf8 ("the diabolical gene" per Quay) in his testimony.
Erdman even interviewed Quay for his podcast back in November 2023. It appears to be very likely that Erdman is the same person who was the whistleblower to the House select subcommittee in September 2023. It's unlikely Erdman had any relevant information to report at the time. He's said on multiple other occasions that he was unemployed circa late 2022 and beyond. His report now only discusses things he learned via work for Gabbard's investigation. If that's the case, Senators Paul and Johnson were very nice to give Erdman the opportunity to walk back false statements about bribes to analysts and Fauci secretly visiting the CIA.
Oh, and Erdman and his organization also settled a case with DOJ last year with an undisclosed windfall to benefit the organization (Feds For Medical Freedom v Biden). This was a case that was going nowhere since the federal vaccine mandate had long ended.
But no one's going to say this is some sort of conspiracy in which Trump's DOJ pays off Erdman in exchange for Erdman testifying to vague misdeeds at CIA under Biden. Because all it is is a dumb waste of money by an incompetent government and has nothing to do with how the pandemic started.
Forget COVID origins, what about Hantavirus origins?!
@bens Conspiracy theorists' heads are spinning now deciding between focusing on:
CIA whistleblower who in reality revealed that access of Gabbard's team to all of the intel community's reports failed to find any cover up. He did find that there was consensus in the intel community against some specifics of various popular COVID origins conspiracy theories (ie origin in November rules out September database blah blah blah), but of course people just ignore that. The fact that he revealed nothing about suppressed analysis and/or evidence is proven by the fact that everyone's focused on a bullet point in his testimony that says Fauci did something "intentionally" on the other side of the world.
Hantavirus outbreak origins
Ebola outbreak origins (they'll get around to it if they haven't yet; surely it'll be unusual in some way that can be said to be suspicious)
The conspiracy behind firing Tracy Beth Høeg and Marty Makary. Apparently it's impossible to realize two people simply had zero aptitude for the jobs they were given and did bad jobs, irrespective of what the administration's priorities are.
@MachiNi I left out African Swine Fever Virus that also came up again today as a possible recent lab leak.
@MachiNi not until they started impacting domestic policy and international relations. It’s fascinating how stuff that’s obviously false is impacting the real world.
Take Robert Kadlec. He thinks SARS-CoV-2 was a bioweapon. He’s assistant secretary of defense in charge of policy for a portfolio combining nuclear, chemical, and bio defense.
Unsurprisingly since USA doesn’t have offensive chemical and bio weapons capabilities, creating this position led to the mission creep to be able to credibly threaten to use nuclear deterrence to respond to non-nuclear unconventional attacks. This requires being able to attribute the attacks (also a challenge for some nuclear scenarios).
The guy in charge of this wants a system that would’ve determined SARS2 was a bioweapon! Albeit, not one intentionally deployed so probably no nuclear response, in that instance.
So, having the entirety of the US government and many other governments bewildered by conspiracy theories because it’s politically problematic to loudly call bullshit increases the likelihood of dumb responses to future rare events. Even aliens are taken too seriously because it’s a pet issue for a couple folks in Congress. But perhaps that’s a net positive for libertarians who think there’s a negative opportunity cost to government spending time on real things 😂
Not sure exactly why this situation exists; some parallels with 2002/3 but also different in many ways.
I don’t have as much insight into China, but perhaps it’s not so different there. Their white paper on COVID origins is quite similar to the Covid dot gov page. Answering the origins question isn’t a huge deal; I’m concerned with this irrationality coming up again the next time there’s a similar question.
Anyone have a guess at who Rand Paul's whistleblower will be at the hearing tomorrow? Check out his posts on X on his two accounts the last few days if you want to see some potential hints.
If someone wants to make a separate market about whether or not the whistleblower will offer evidence that objectively increases support for a laboratory origin of COVID-19, I might bet on that.
The hearing should be shown live here tomorrow: https://www.congress.gov/event/119th-congress/senate-event/338388
@zcoli https://nypost.com/2026/05/12/us-news/rand-paul-says-cia-whistleblower-will-allege-ongoing-deep-state-coverup-of-covid-origins-at-public-hearing/
This story has the most details I've seen so far. They don't exactly match the details Paul gave in a different interview I saw, but that's to be expected given this is all quite likely to be as factual as David Grusch's testimony was. Not sure if other dimensions will be invoked to explain the absence of evidence to back allegations this time around, though.
Very willing to eat my words if the guy shows up with evidence that the CIA suppressed actual evidence of a lab leak. But I figure it's the same guy who previously blew the whistle about Fauci's secret trips to Langley and the CIA paying off analysts to conclude natural origins. Those were never substantiated even though some in Congress took them seriously enough to investigate.
@MachiNi someone duping Paul with a whistleblower disclosure that can be read in one of two ways but is guaranteed to be read one way by conspiracy theorists in order to testify before the committee makes for a nice fiction to think about though.
Did anyone read Matt Ridley's paper that he says is too hot for peer review?
Is this really the best argument that lab leak advocates can come up with? It appears to have won approval from all of the lab leak advocates, so I guess so. But, does Ridley know what he's talking about?
Let's take Reference 33, for example, which Ridley uses as evidence that phylogenetic analysis dates the first infection to "the end of September—beginning of October."
First, it's weird he doesn't just read from the abstract which says they "estimate the time of SARS-CoV-2 zoonotic spillover between August and early October 2019." He picks a subset of the analysis in second.
More importantly, the analysis is very, very obviously invalid. Let's just take a look at the first time in the paper that one of those estimates (August 2019) pops up and see the corresponding figure (Figure 3A):
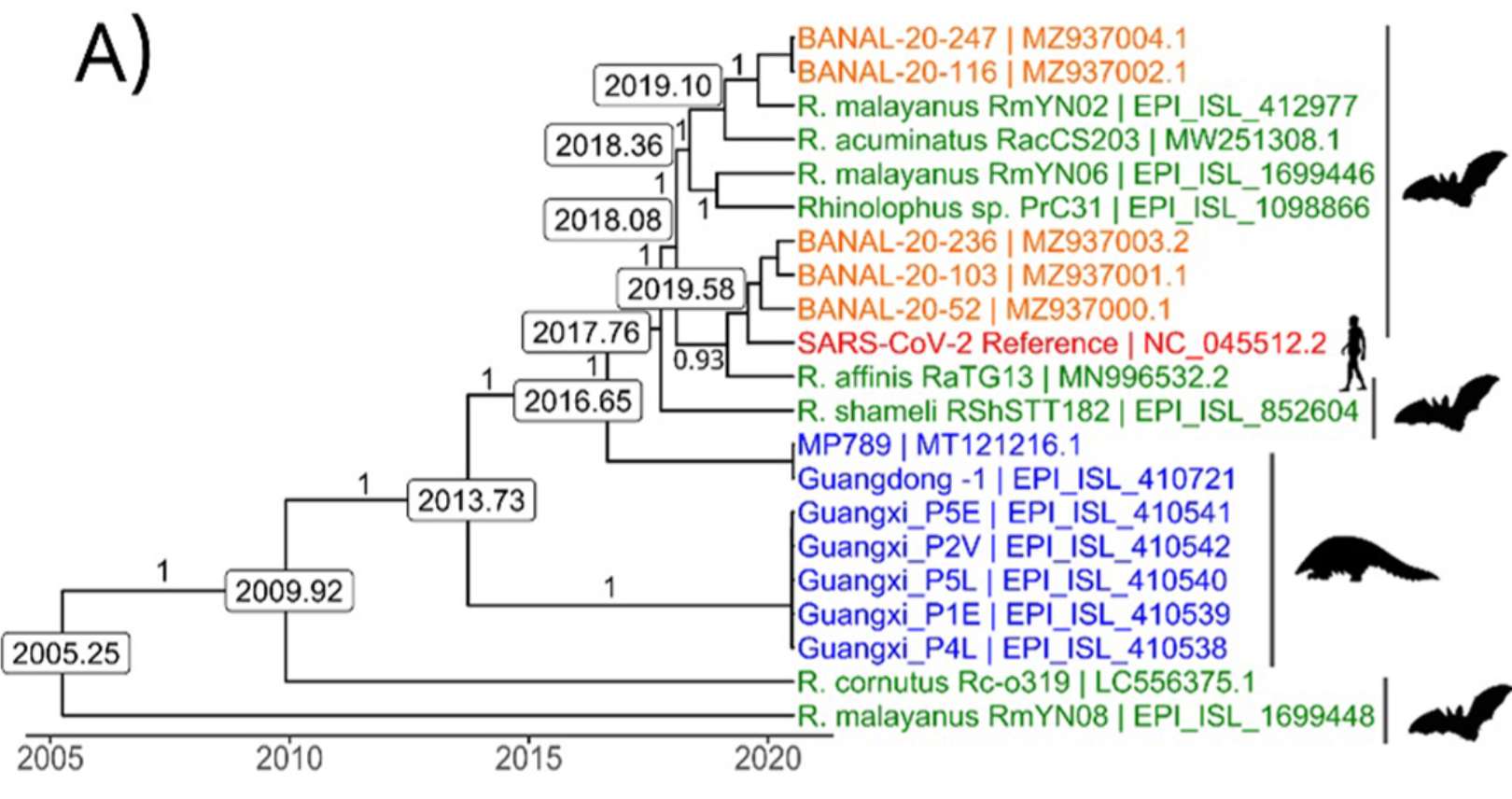
In case you are unaware, the "13" in RaTG13 stands for 2013, the time it was sampled.
In this figure, the authors show that they estimate that the time the common ancestor of RaTG13 and SARS-CoV-2 was around was in 2019. "Lab leak or not?" is a somewhat important question, but Ridley is missing the even bigger time travel story here!
I didn't read the whole thing but digging into a random citation I see an immediate error:
This is underlined by the fact that the WIV’s bat and rodent pathogen database, with a password-protected section for unpublished viruses, was made inaccessible to outside access on 12 September 2019[63], prior to the outbreak, and has never been accessible since.
Cite 63 is https://www.help.senate.gov/imo/media/doc/report_an_analysis_of_the_origins_of_covid-19_102722.pdf
Which actually says:
On September 12, 2019 between the hours of 2:00 and 3:00 a.m. local time,166 the WIV took down its online depository of data on viral sequences called the Wildlife-Borne Viral Pathogen Database.167 The database was intermittently accessible from December 2019 to February 2020, before being permanently taken offline February 2020.
This error is, "no doubt", accidental.
@draaglom yeah I’m not intending one example to be exhaustive.
The one and only figure in the paper being a screenshot complete with capturing a blinking cursor and some red squigglies for spell check speaks for itself.
Ridley says it went out for peer review — amazing it didn’t get desk rejected.
A funny thing about the databases (there are many databases in the conspiracy theory) is that lab leak people don’t notice when most of them went online again years ago.
https://econjwatch.org/articles/an-article-in-science-on-covid-origins-contains-a-fundamental-error. An Article in Science on Covid Origins Contains a Fundamental Error
@George If anyone knows someone lucky enough to have not really heard much about the “lab leak” question and wants to try this experiment, I’d be really interested in the results:
Print out this paper and lock your subject in a room for two hours without their phone.
Their task: thoroughly review the paper and identify what the major scientific question is that it addresses is, to list the evidence important to answering that question, and to determine Weissman’s conclusion(s) and rationale.
When their time is up, ask them whether Weissman’s POV is that COVID-19 spread from the Huanan market. If they say they don’t know, give another 15 minutes to review to focus on and answer that question.
Once they answer “no that’s not his POV”, ask what the missing evidence is. They’ll say that the missing evidence is that Lineage A wasn’t in Huanan market.
Say, “What do you mean? Back in February 2022 there were three famous manuscripts published almost simultaneously. One of them reported that Lineage A wasn’t literally found inside of Huanan market! Didn’t Weissman discuss this?”
Give another 15 minutes to review the manuscript if it’s required to find that the answer is “no”.
Tell the person that the fact that Lineage A was found in Huanan market is discussed prominently in many of the papers referenced by Weissman, and Weissman has discussed this at length for years elsewhere.
Finally, ask them to summarize their feelings about the manuscript in 50 words or less and report the results back in this thread.
As a follow-up if you’ve gotten their attention, provide the preprint for Pekar 2022 that contributed to the news coverage on this question, and ask them to find the “math error” that Weissman complained about and to see if, in fact, Weissman falsified the whole paper like he claims.
Because it’s absurd that Weissman is claiming to have overturned an entire paper because he disputes one part of it that’s not its singular main result and arguably not even a main result (it wasn’t in the preprint; added during review).
And he’s not disputing a “math error” — he’s demanding answering a question that Pekar 2022 plainly did not claim to answer quantitatively. For sure, like basically all complex scientific manuscripts, especially in journals like Science targeting general audiences, the paragraph he’s complaining about could’ve been written more precisely. No argument there from me. The paragraph also should’ve kept the “clock reversal” part that was in that section of the preprint, rather than moving it elsewhere during editing.
But there’s zero doubt that Weissman is comparing different hypotheses than Pekar 2022:
The relevant phrase: “… we inferred substantial support favoring separate introductions of lineages A and B (BF = 4.3 …”
This was clearly conditioned, for better or worse, on comparing (1) separate introductions of A and B to (2) a single introduction of A, B, or an intermediate between the two.
It was clearly about comparing phylogenetic topologies from a single introduction sampled once or twice. Nowhere does P2022 claim to simulate multiple introduction. Not considering relative dynamics of the two introductions; just the ultimate topology.
The paper never concludes “two and only two” introductions; always “at least two”. The paper specifically says introductions of identical genotypes aren’t distinguishable, for example.
The conclusion of “at least two” is a subjective conclusion arising from two observations: (1) relative rarity of a single introduction producing observed topology compared to a specific multi introduction scenario, (2) clock reversal — lineage A is the likely common ancestor, yet lineage A looks “newer” with less divergence than lineage B, and less lineage A than lineage B in the world.
Weissman compares hypotheses of 1 vs 2 introductions of any sequence; clearly not what P2022 did for any reader paying attention.
Weissman extends P2022 to add an unrealistic model of phylodyanmics of two and only two spillovers. This isn’t correcting a P2022 error; it’s doing something totally new.
Weissman ignores “clock reversal” entirely.
Weissman also badly misrepresents many of the papers he cites. To the extent where it’s doubtful that he’s ever read them and suggests that he’s repeating things he read on Twitter rather than reading the literature.
His statement that Worobey 2022 investigated “superspreading” from Huanan market when both Worobey 2022 and the very first thing Weissman cites (a NYT article) say literally the opposite is certainly interesting.
He states that the most likely ancestor was lineage A, citing a bunch of papers that give no quantitative likelihood in support. The first paper to adequately estimate this qualitatively: Pekar 2022! Weissman doesn’t cite it on this point. That was one of its main results! More important than BF=4.3. Weissman appears to be unaware of this aspect of the very paper he’s claiming to have entirely falsified.
Lastly, Weissman implicitly incorporates his substack post by citation. His post remains full of errors despite years of removing very obvious ones (e.g., an estimate of spillover date from a figure which the common ancestor of RaTG13 and SARS-CoV-2 existed well after 2013… this is impossible). The main piece of evidence he focuses on is some molecular biology creationism about restriction sites that would embarrass creationists and was proven false back in September-October 2022.
https://videocast.nih.gov/watch/244438a5-0e6b-11f1-9f14-124f0a52e769
NIH Scientific Freedom Lecture – Viral: The Search for the Origin of COVID-19
@George Ridley kicks off his summary:
To summarize, ladies and gentlemen, in 2017-18 in Wuhan and nowhere else, the Wuhan Institute of Virology shifted their priority to 20% divergent viruses from SARS…
I assume he leads with his strongest argument. So, if that falls, the rest is likely to fall apart as well.
So, does Ridley have a point? Is SARS-CoV-2, a bit over 20% different from SARS spike, eerily similar to what researchers in an international collaboration including WIV were interested in looking for?
I decided to take a look using a deduplicated set of spike sequences from SARS-like viruses.
Nope. Just about everything is about 20% different.
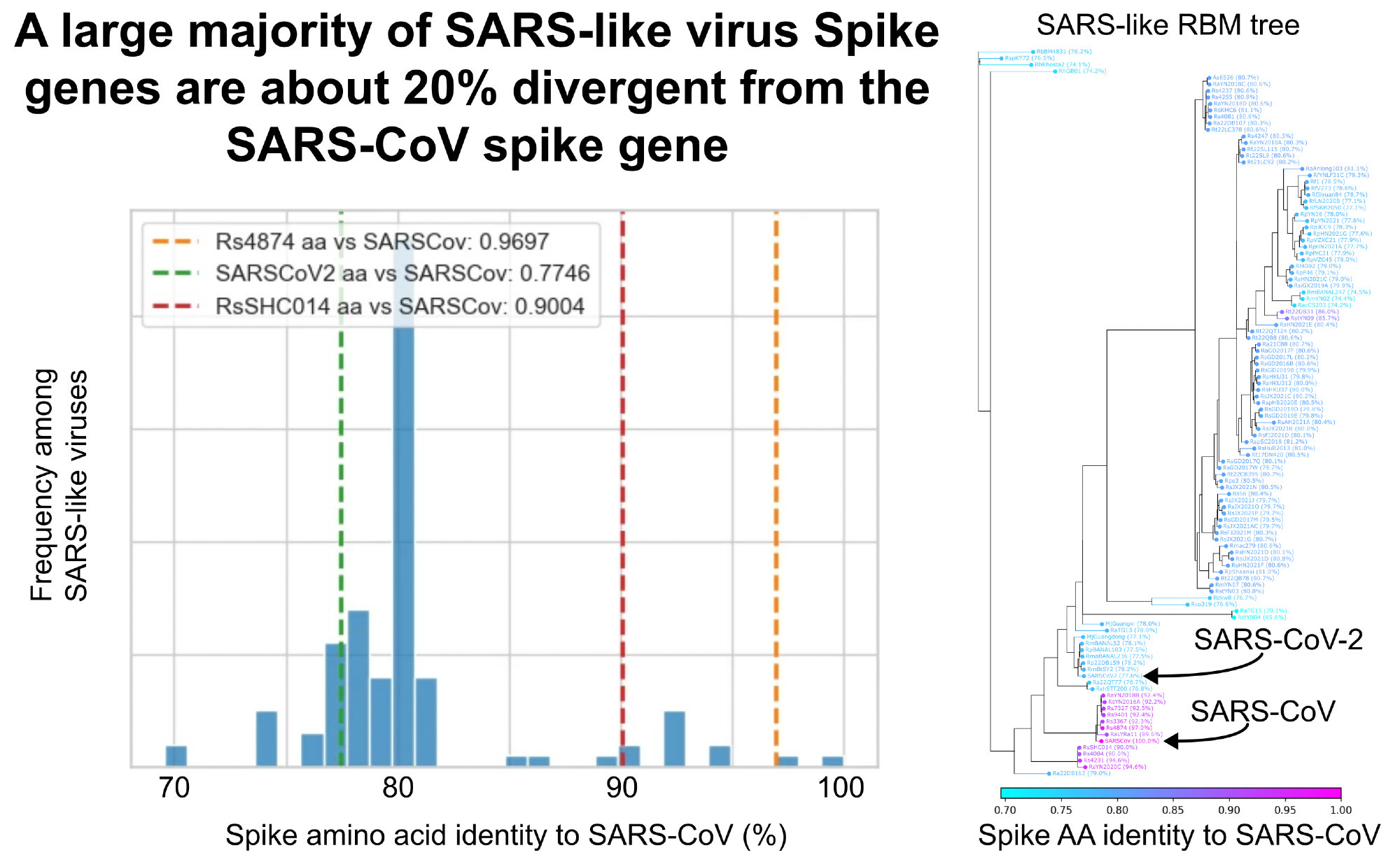
Did Ralph Baric at UNC Create SARS-CoV-2?
@George Brownstone is gonna drive someone crazy enough to want to start a pandemic to end the world with that web design choice to read the article to you if you click anywhere.
So I didn't read the article... I just looked at the source and saw "utm_source=chatgpt.com" for the link to Sachs' own article.
So... Sachs is outsourcing analysis of his own article to ChatGTP?
